If diversity is not used, why conserve it?
By offering more effective access to crop diversity and information, the Platform contributes to increases in the productivity, resilience and nutritional quality of crops.
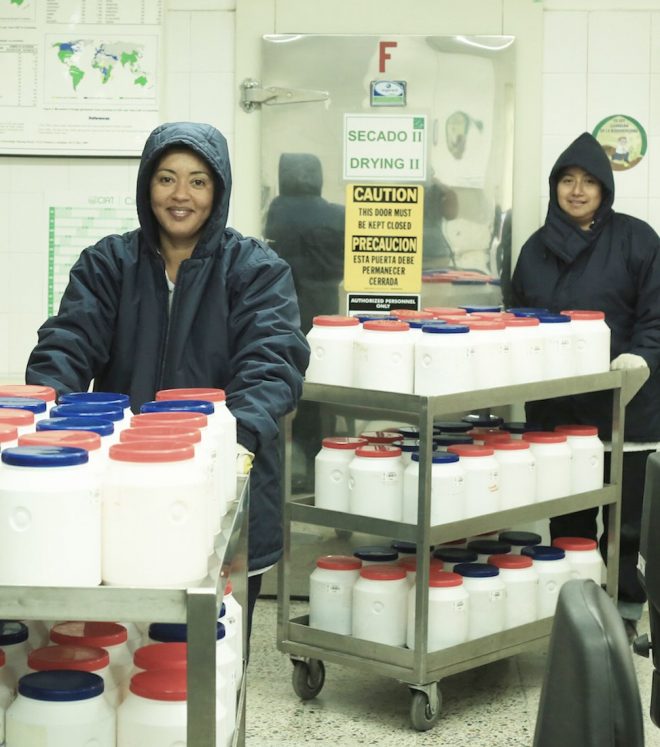

Credit: Shawn Landersz
The Use module goes beyond increasing the number of accessions distributed. It tackles some fundamental constraints that genebank clients like breeders face when selecting crop diversity – helping them to make better informed and more precise choices, uncovering new traits or characteristics in well-known crops or just making it easier to get hold of clean materials rapidly. The Use module addresses all these aspects.
The Use module works closely with the CGIAR Research Initiatives – major users of crop diversity – to ensure that breeders and other researchers are closely connected with the genebanks.
The goal is to enable more effective access to and use of crop diversity, by all who might need it.
Selection of material
The biggest challenge breeders face is to select parents that contain the traits they are looking for. Sometimes they make do with a random subset of accessions, screening them all to find the likely contenders. If the trait is rare, difficult, slow or expensive to assess, the breeder may not be able to screen sufficient accessions to find it. Even when rapid, large-scale screening is possible, a trait may remain undiscovered because it is not expressed or it may require a specific environment. Again, the breeder faces failure.
The path to success involves more – and more useful – information about each accession. Genesys is an information system being developed with the Use module to give researchers an effective way to search for information about genebank accessions and then order samples directly from the genebanks. The Use module helps bring new sorts of information into Genesys. For example, knowing the environments in which accessions were originally found can enable predictions of what stresses they are likely to be adapted to, including those associated with climate change.
The Use module aims to take approaches such as Focused Identification of Germplasm Strategy (FIGS), originally championed by ICARDA, and makes them available to all genebanks and their users.
Similarly, genebank staff routinely score accessions using traits that are highly heritable and little affected by environmental interactions. The Use module helps CGIAR genebanks to bring these characterisation data into Genesys to let researchers query the data and build their own subsets of accessions to screen.
Perhaps the most relevant data for breeders, however, is collected not by genebank staff but by the breeders and other users of accessions. These data encompass phenotypic data, gene expression and “-omics” data, including genetic sequences. Together, such information offers a much clearer insight into the likely performance of specific accessions, a huge resource to improve the use of genebank material. To date, however, much of this information has not been tied directly to specific accessions and in any case has not been available to genebanks to share with other potential users. The Excellence in Breeding Platform is developing systems to facilitate data generation and sharing of this nature. Leveraging a relatively small additional investment, the Use module can build tools that allow genebanks to associate new information with accessions in their collections and dramatically improve the experience of the users in selecting the accessions that they need.
While the Use module works closely with partners in the CGIAR, the tools and services it develops can equally help breeders, researchers, farmers and others outside CGIAR who need access to information about crop diversity and the diversity itself.
Challenging data
One difficulty that has long plagued researchers who generate and use information about genebank accessions is that different institutions often use different data structures. This incompatibility makes it almost impossible to gather disparate sources of information to build a comprehensive picture of an accession’s properties. The Excellence in Breeding Platform is helping to harmonise data gathering by breeders, and the Use module works to ensure that these data are associated with specific genebank accessions.
A second challenge is to track accessions as they move out of genebanks into research plots and beyond. All too often, the genebank’s specific “name” of a unique accession is separated from the accession, making it impossible for information derived from the accession to return to the genebank’s databases. Researchers are beginning to appreciate the need for globally recognised Permanent Unique Identifiers (PUI) and the Use module is promoting the uptake of Digital Object Identifiers (DOI) as the preferred PUI for germplasm samples. A system of PUIs will tie together diverse sources of information and will enable researchers to search across multiple databases for information about accessions, again facilitating the use of genebank material.